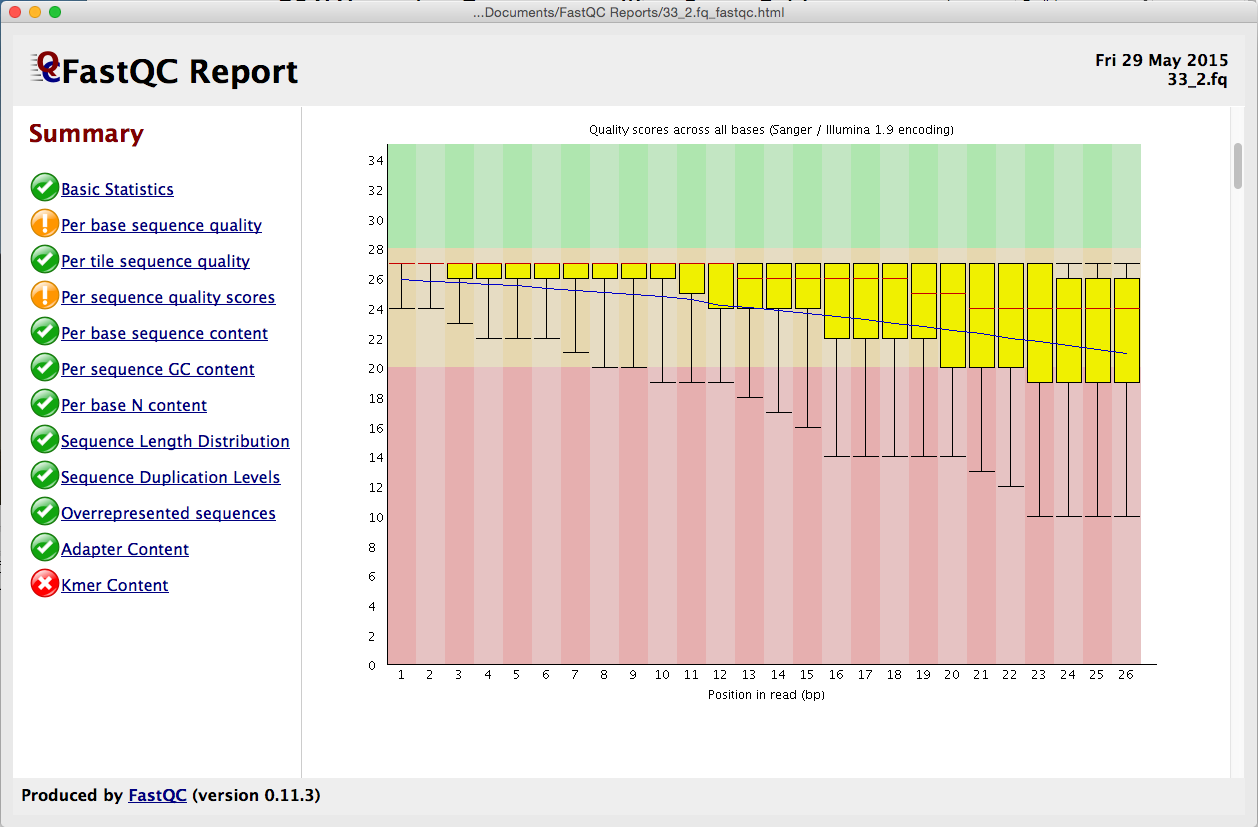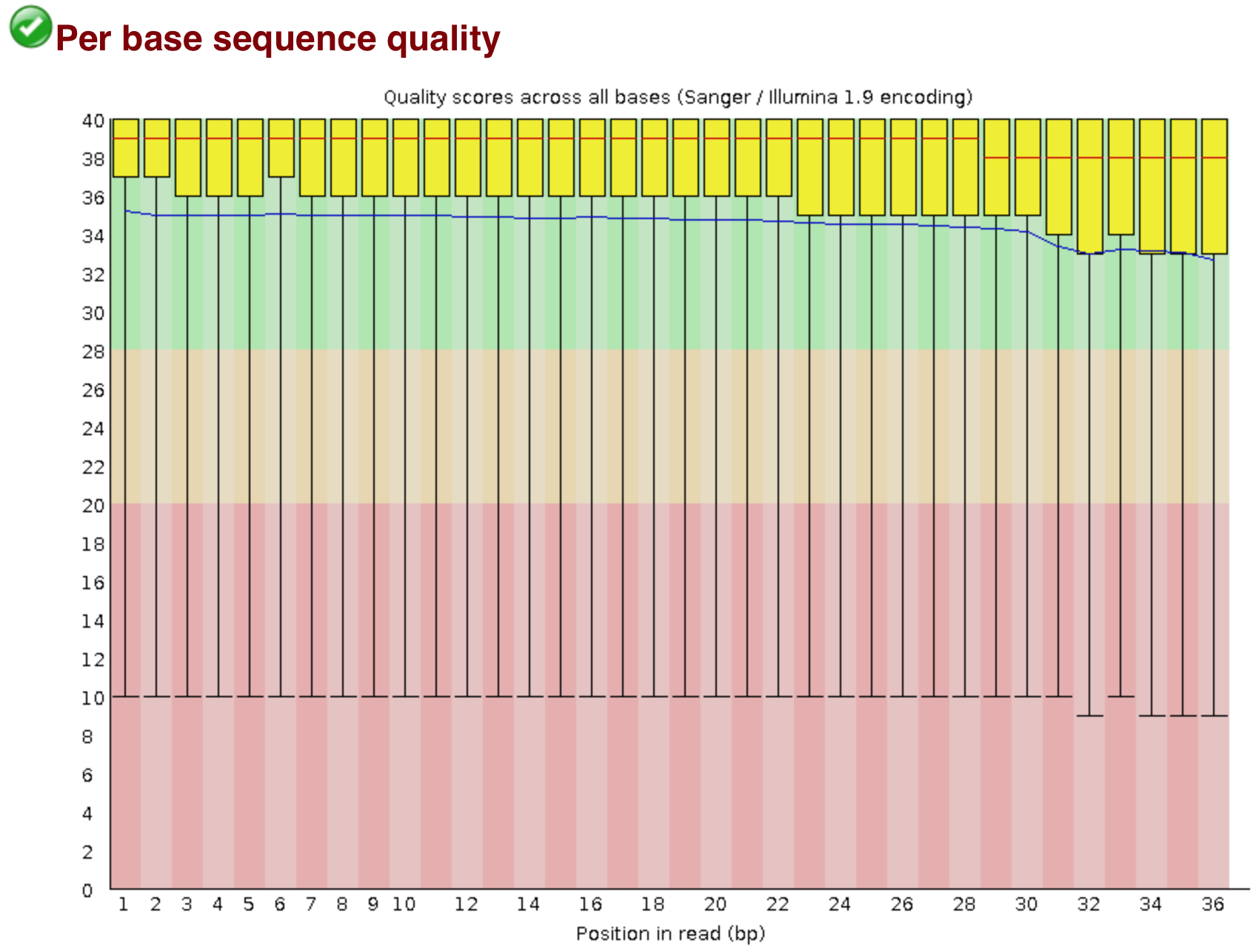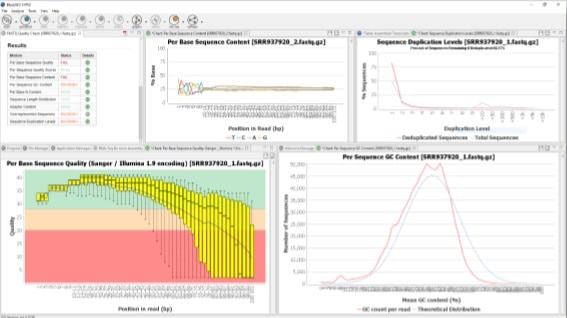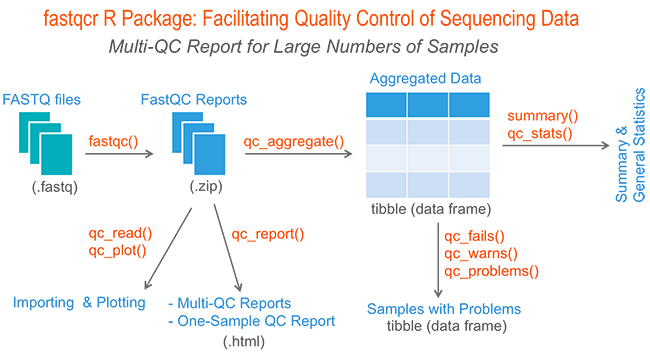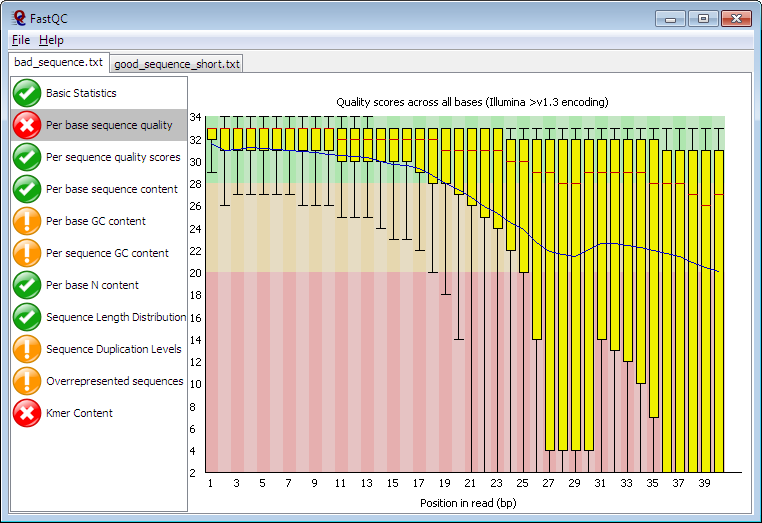
Babraham Bioinformatics - FastQC A Quality Control tool for High Throughput Sequence Data | BibSonomy
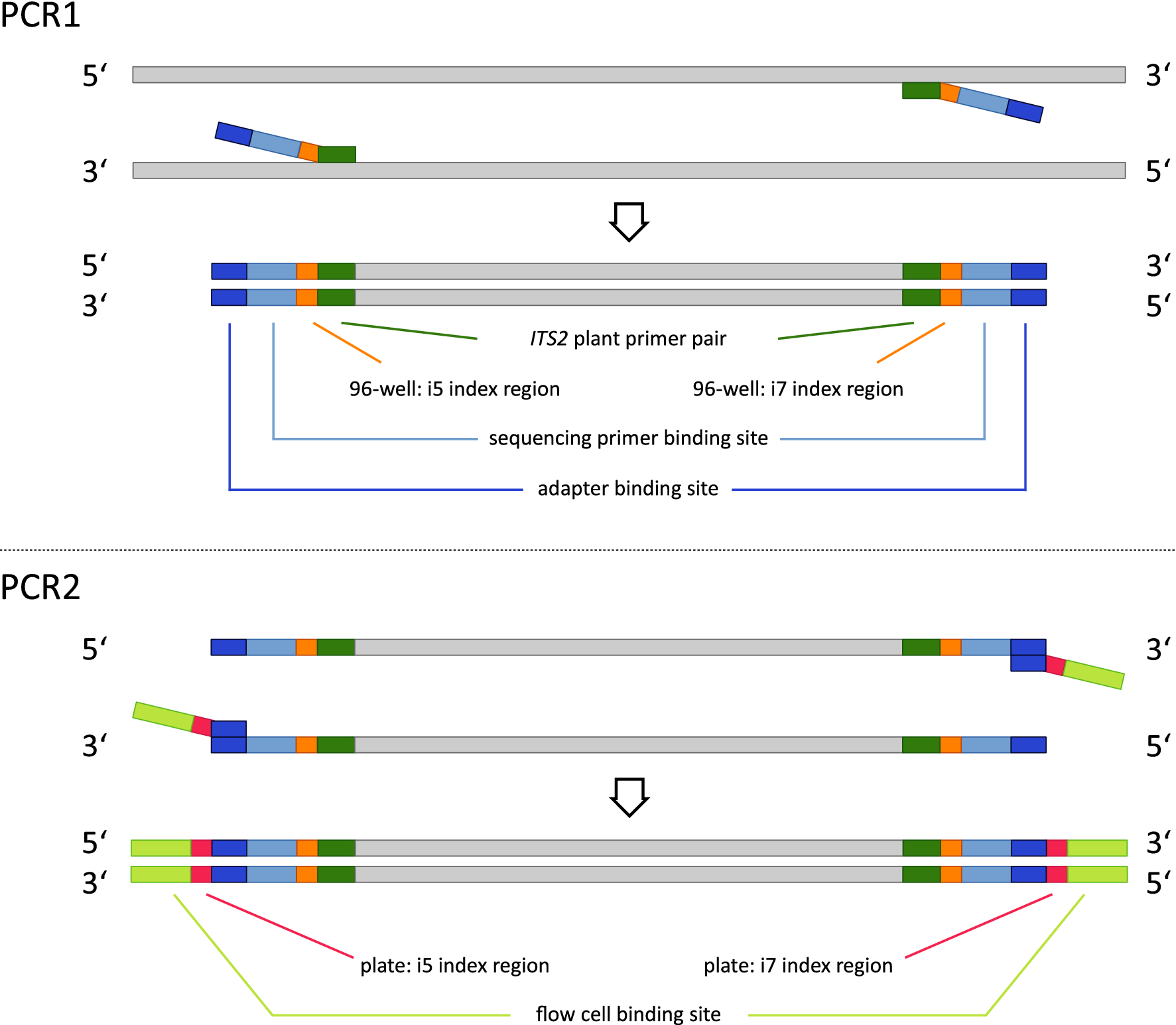
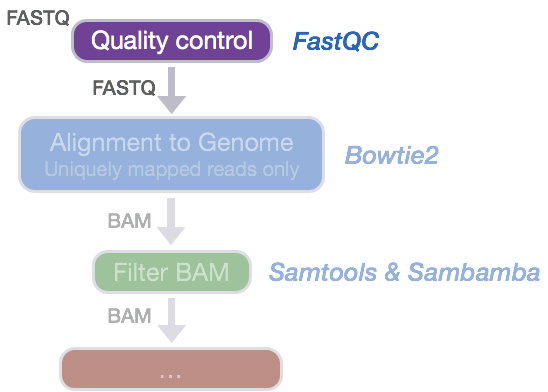
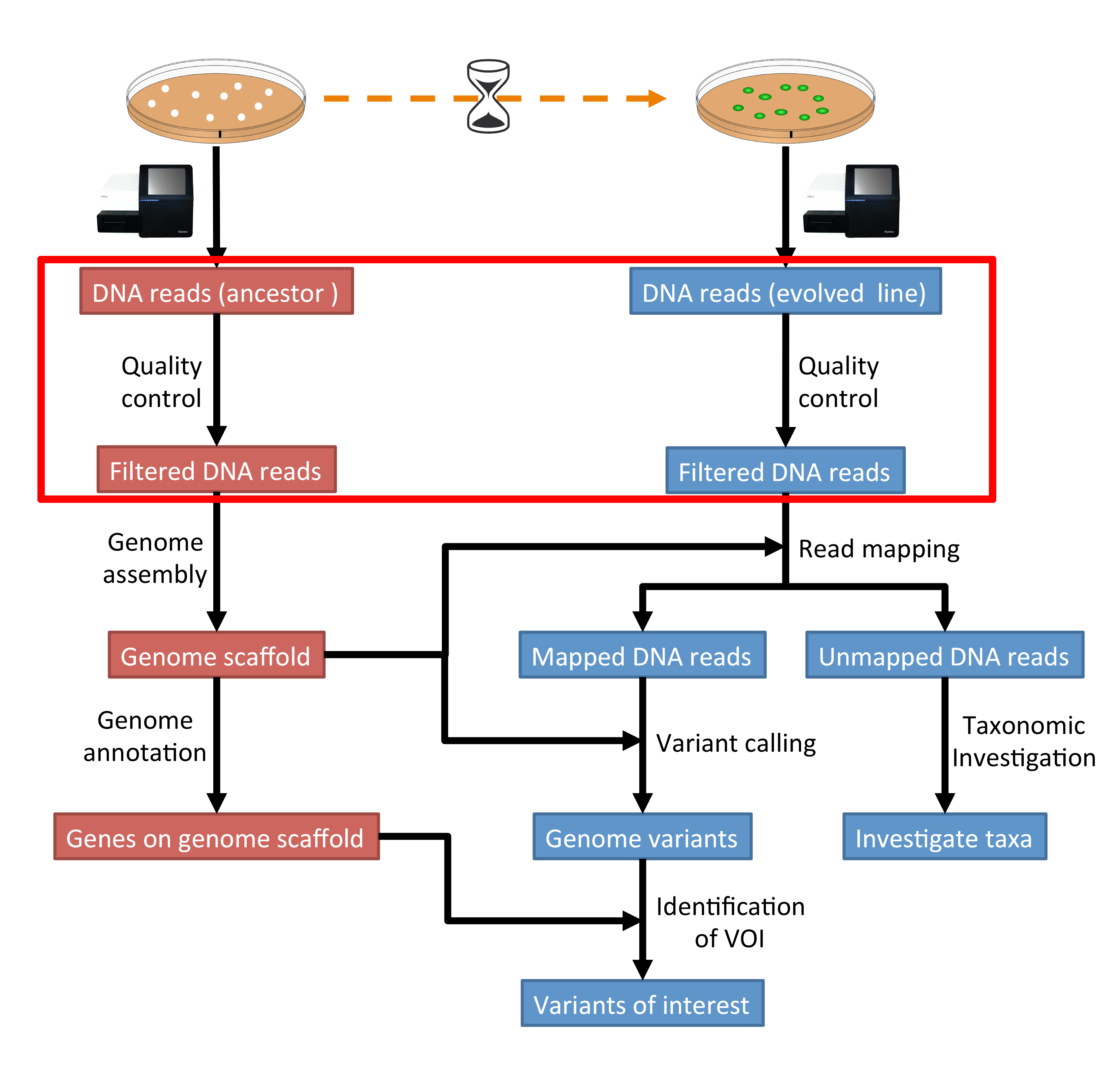
Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports
GitHub - natir/fastQt: FastQC port to Qt5: A quality control tool for high throughput sequence data.

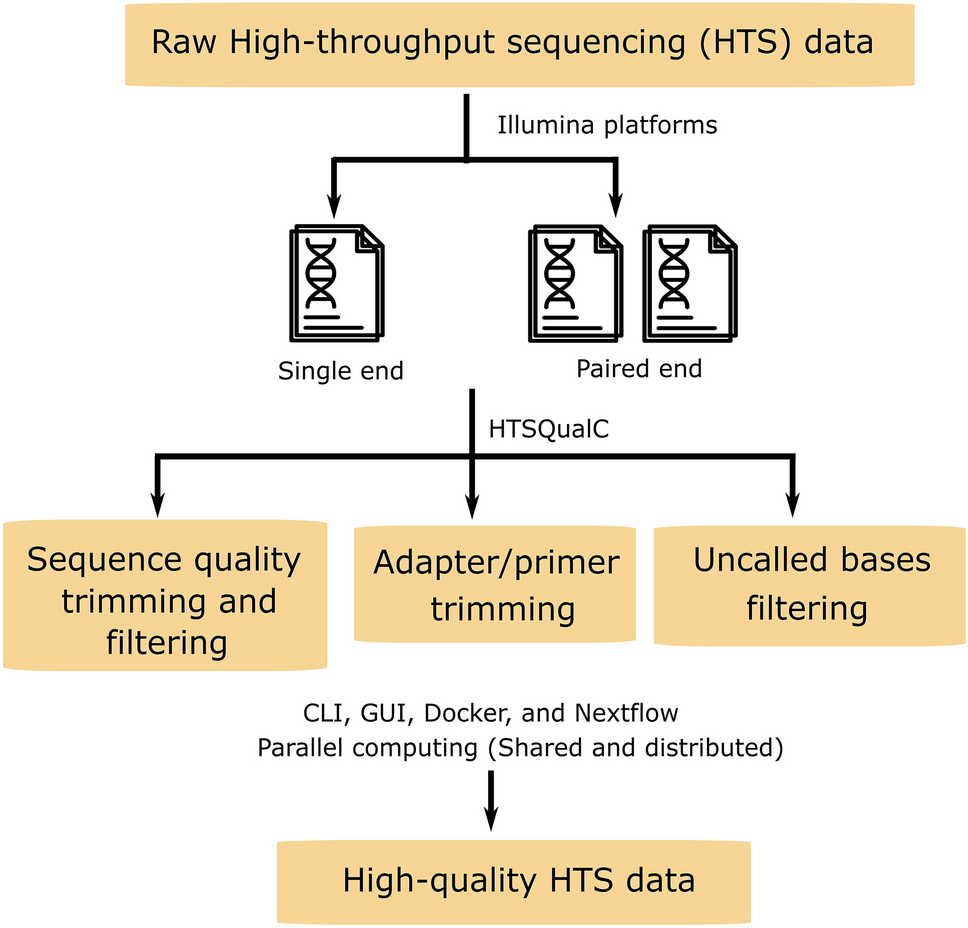
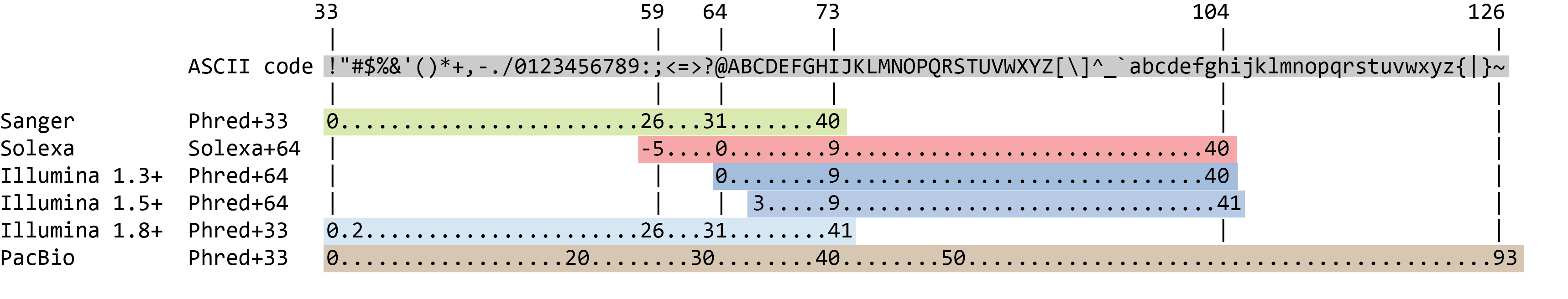
Biomolecules | Free Full-Text | High-Throughput Identification of Adapters in Single-Read Sequencing Data