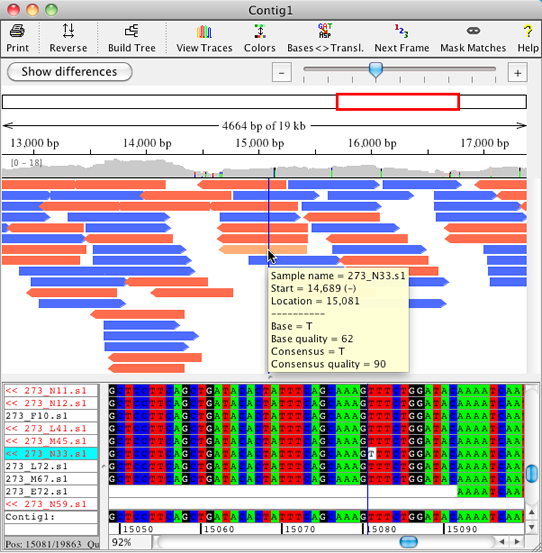
New Features in CodonCode Aligner 4.0 - DNA Sequence Assembly and Alignment Software for Windows and Mac OS X
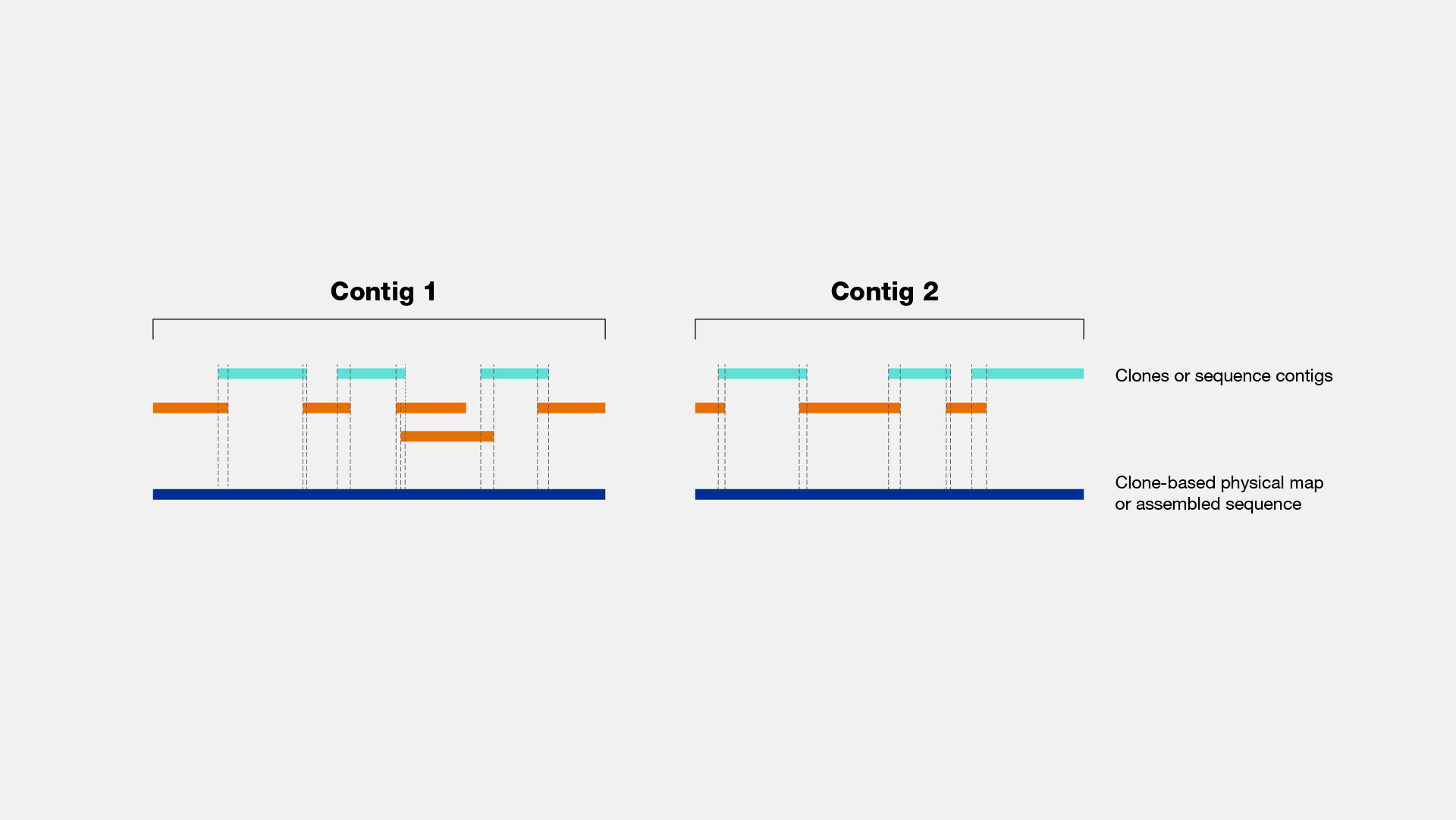
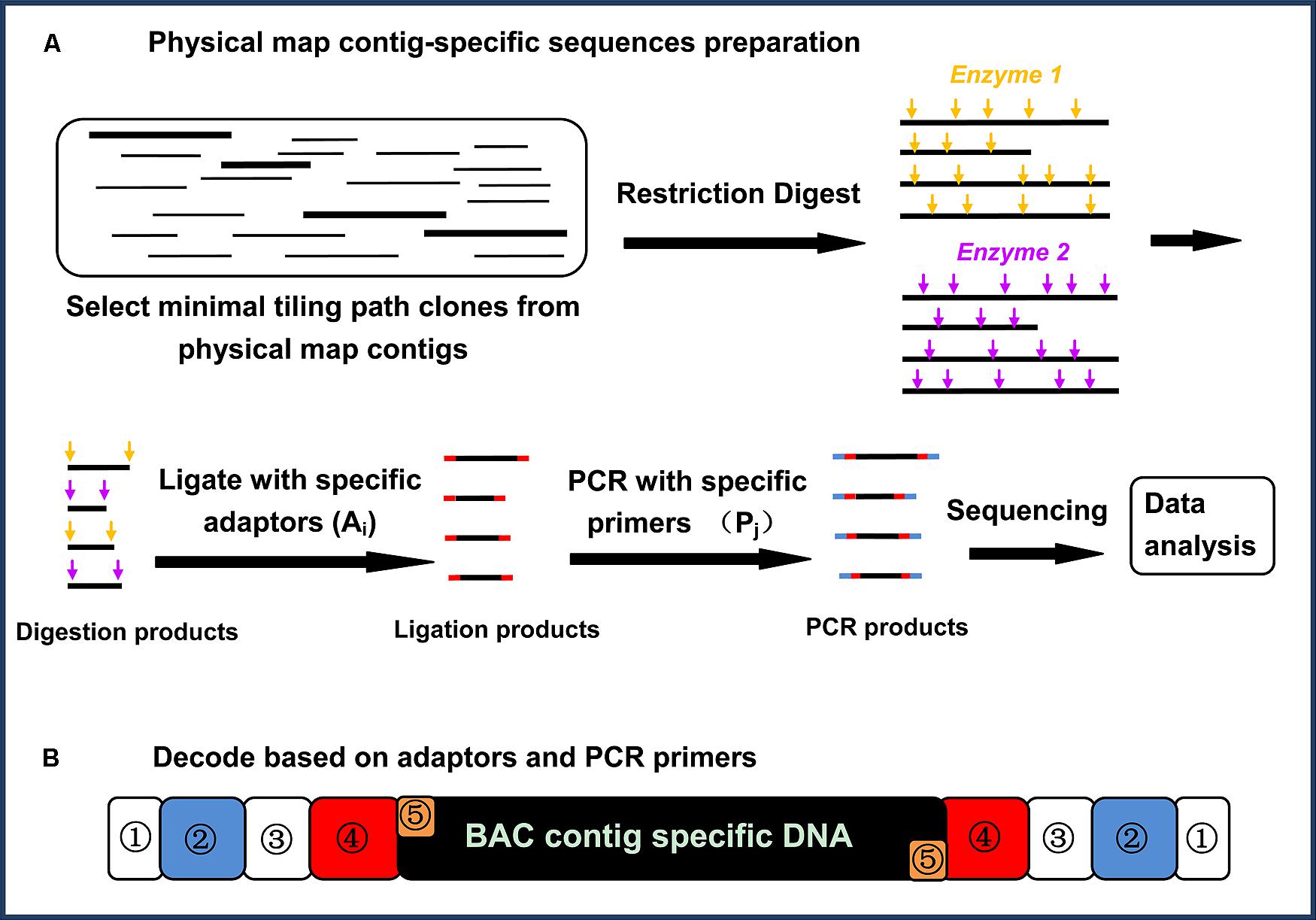
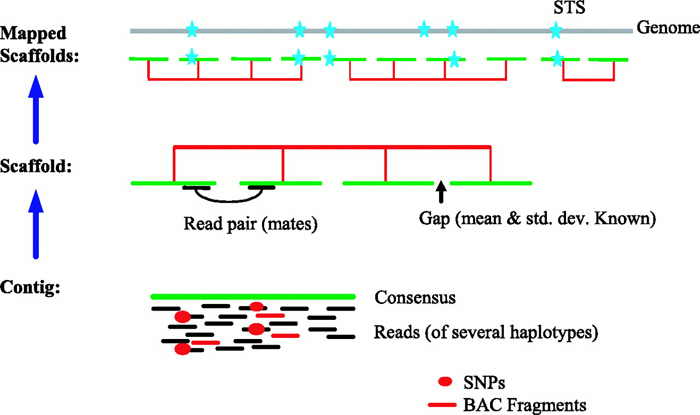
Generation of Physical Map Contig-Specific Sequences Useful for Whole Genome Sequence Scaffolding | PLOS ONE
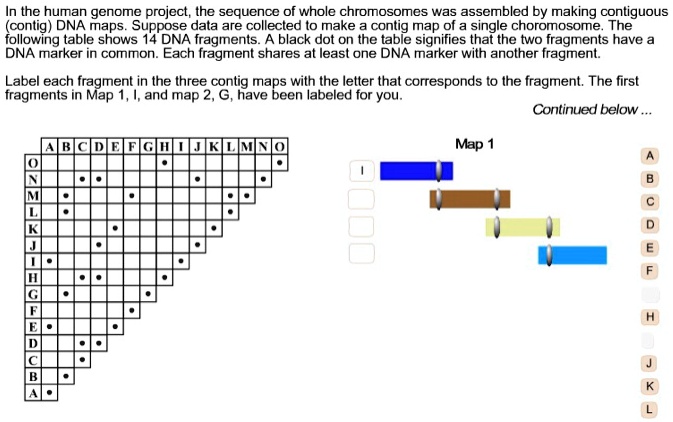
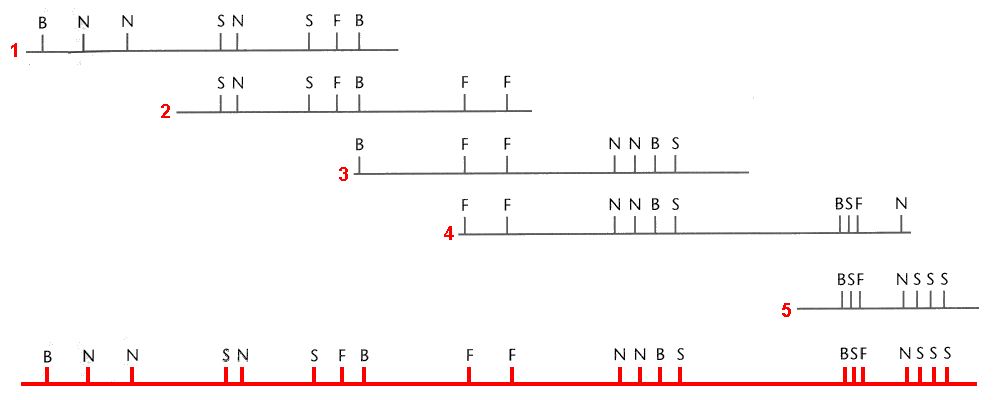
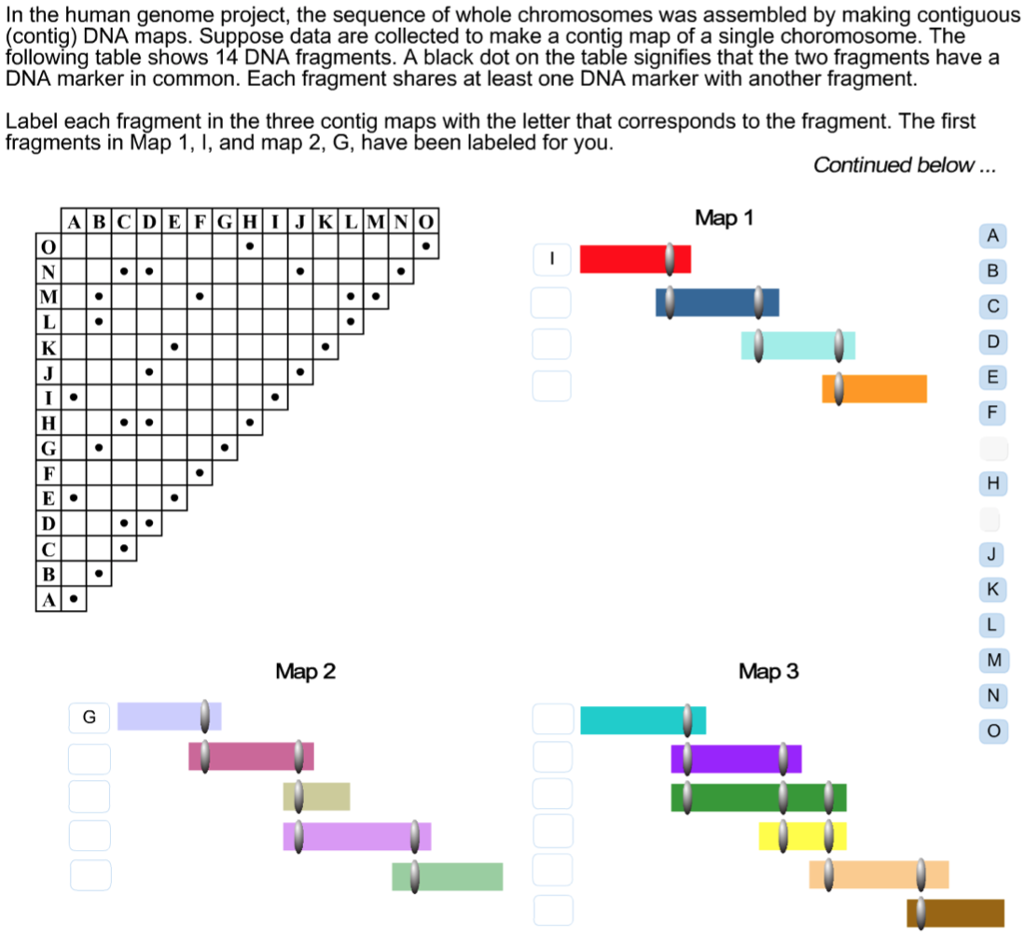
SOLVED: In the human genome project; the sequence of whole chromosomes was assembled by making contiguous contig DNA maps. data are collected make contig map of single choromosome The Suppose following table

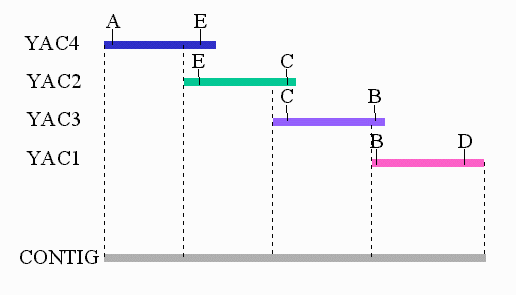
Clone Contig Sequencing Methods | Clone Contig Approach | Whole Genome Sequencing By Clone Contig | - YouTube
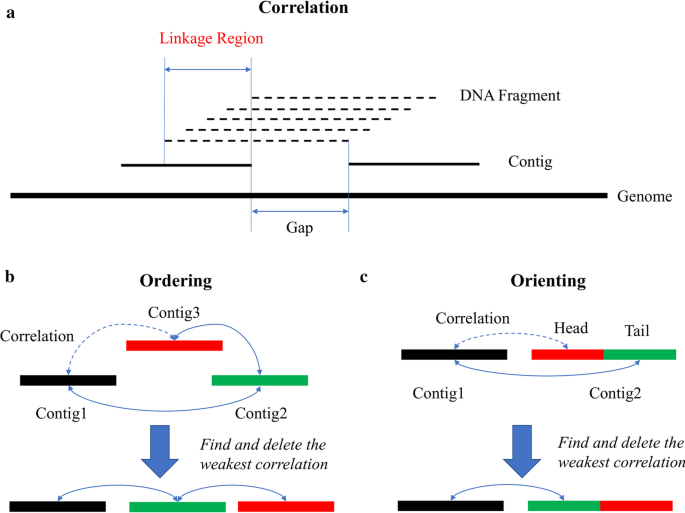
SLR-superscaffolder: a de novo scaffolding tool for synthetic long reads using a top-to-bottom scheme | BMC Bioinformatics | Full Text

Introduction to the principles and methods underlying the recovery of metagenome‐assembled genomes from metagenomic data - Goussarov - 2022 - MicrobiologyOpen - Wiley Online Library

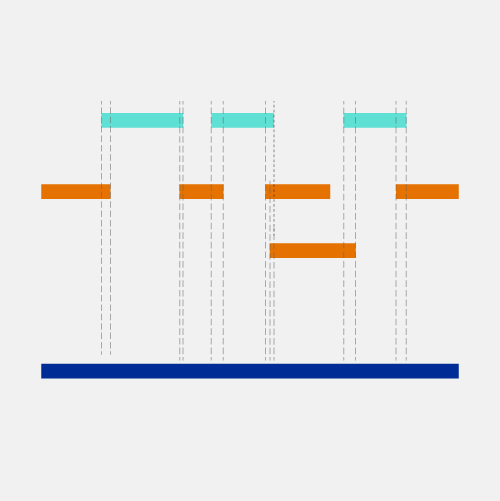
Types of assembled contigs and alignments to REF contigs. a A contig... | Download Scientific Diagram

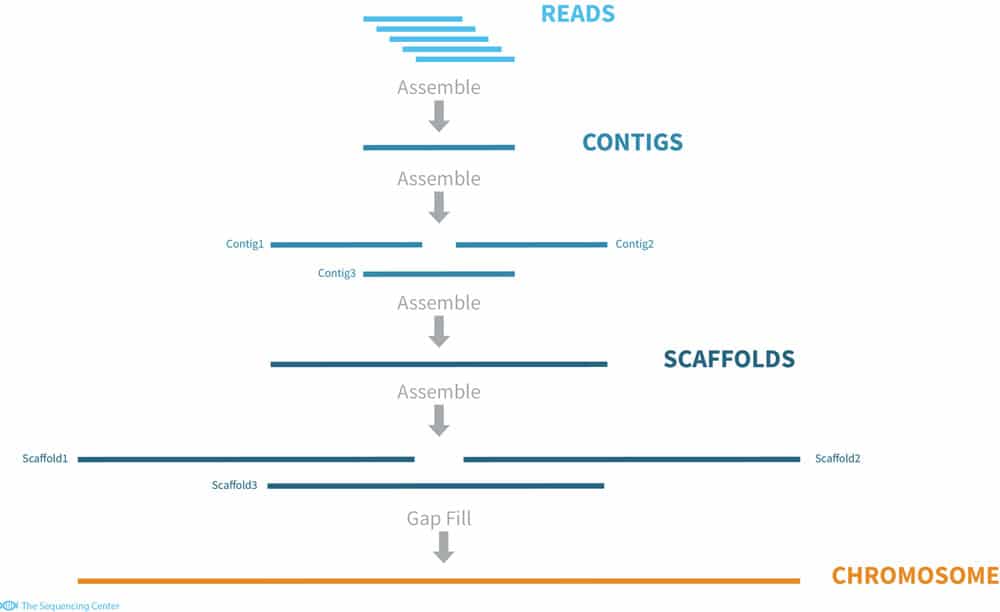
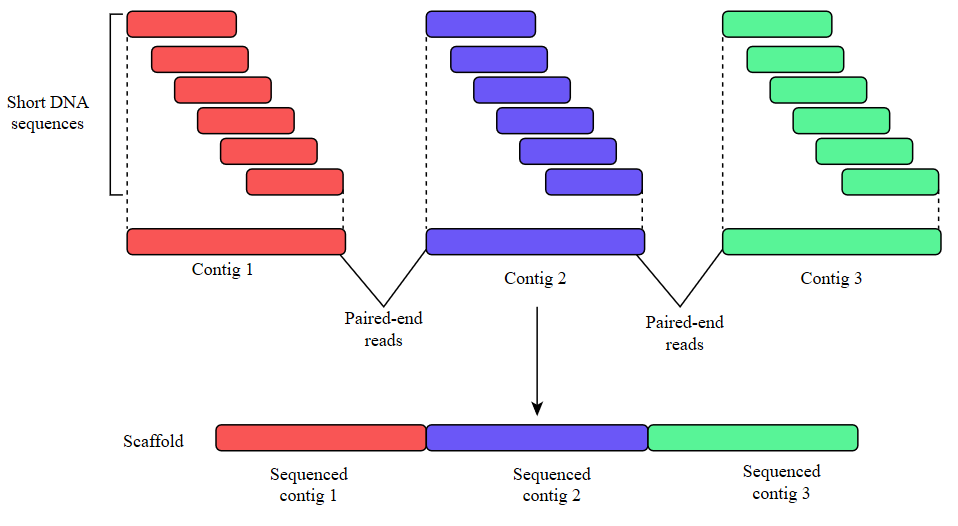
Whole Genome Sequencing Process of Creating Contigs from Fragment Reads | Download Scientific Diagram
![PyDamage: automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly [PeerJ] PyDamage: automated ancient damage identification and estimation for contigs in ancient DNA de novo assembly [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11845/1/fig-1-full.png)